Solution recipes for in vitro electrophysiology from the Miles Lab
Spontaneous Classification
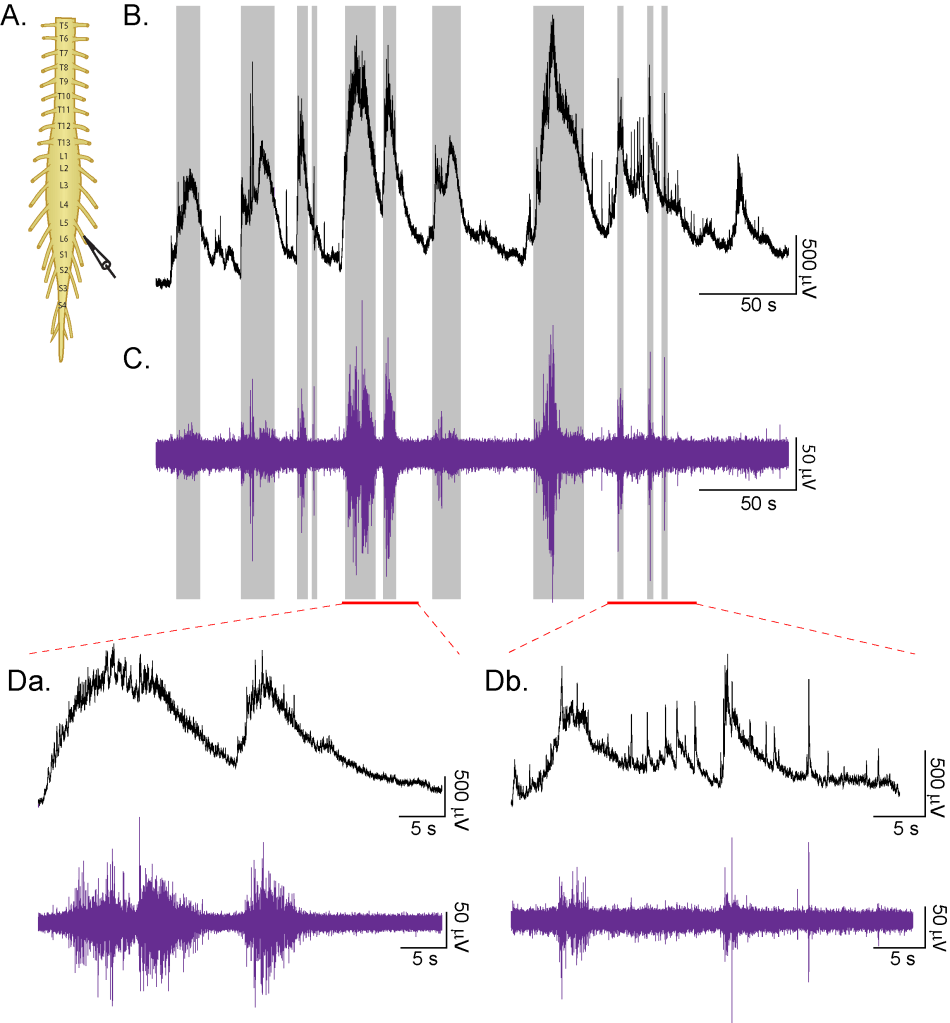
Studies of developing spinal network activity
We developed a software tool in the Whelan Lab, in collaboration with Dr. Ashley Dalrymple (University of Alberta), that uses machine learning to classify can capture the diversity of patterns generated during episodes of spontaneous motor activity recorded from isolated neonatal mouse spinal cords in vitro.
Spontaneous activity is a general feature of developing neural networks and plays a role in circuit formation and refinement.
This tool can be used to study plasticity of developing neural networks.
The custom MATLAB code for this tool, called SpontaneousClassification, is freely available for download in an open-access data repository as an executable file, along with instructions and raw data files used to generate this tool.
The report generated from this work can be found in the following paper published in the Journal of Neurophysiology.